A basic premise of cladist systematics is that evolutionary novelties (derived character states, characteristics of a taxon that differ from the ancestral form) reveal phylogenetic relationships. Such novelties are called apomorphies; synapomorphies are when two or more taxa share such a derived state. On the other hand, shared primitive character states, called plesiomorphies, reveal nothing about the relationships of taxa at the level they are being studied since the symplesiomorphies (primitive character states that are shared) have been inherited from the ancestors of all the taxa. Look at the monophyletic part of the figure below, and assume fur is a plesiomorphic character that has been evolutionarily lost in B and E. The fact that A, C, and D have fur cannot be used to infer a closer relationship among them than with B or E.
A foundation of cladistics is the premise that new character states arise at the time of speciation (for practical purposes, most schools of cladistic thought also require that the parent species ends at the time of speciation, leaving two or more daughter species). [However, keep in mind that cladistics is not a monolithic discipline, and a number of varieties exist.]
Necessary concepts here (and elsewhere in systematics) are monophyly, paraphyly, and polyphyly.
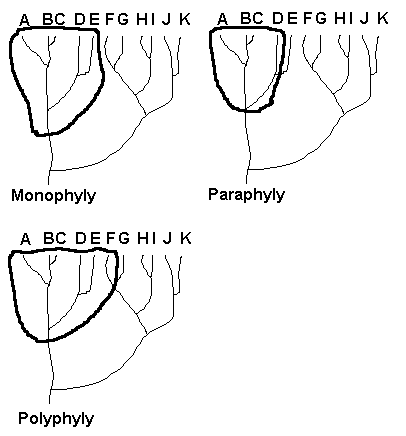 Diagrams showing example of mono-, para-, and polyphyly. The bold line encircles
members considered to belong to the taxon under consideration. In monophyly, all members (A-E) of the taxon share the most recent common ancestor.
In cladistics, strict monophyly is required in that all members descended from the common ancestor must be included in the taxon; in evolutionary classification, some descendents may be
segregated out as separate taxa (e.g., class Aves and class Mammalia from class Reptilia). In the paraphyly example, taxon E is omitted from the higher-level taxon, making that higher lever taxon
paraphyletic. In the polyphyly example, taxon F, which does not share the most recent common ancestor with the others, is included in the higher-level taxon. Under cladistics, only strict
monophyletic taxa are allowed and, in fact, the whole methodology is aimed at discovering monophyletic taxa.
Diagrams showing example of mono-, para-, and polyphyly. The bold line encircles
members considered to belong to the taxon under consideration. In monophyly, all members (A-E) of the taxon share the most recent common ancestor.
In cladistics, strict monophyly is required in that all members descended from the common ancestor must be included in the taxon; in evolutionary classification, some descendents may be
segregated out as separate taxa (e.g., class Aves and class Mammalia from class Reptilia). In the paraphyly example, taxon E is omitted from the higher-level taxon, making that higher lever taxon
paraphyletic. In the polyphyly example, taxon F, which does not share the most recent common ancestor with the others, is included in the higher-level taxon. Under cladistics, only strict
monophyletic taxa are allowed and, in fact, the whole methodology is aimed at discovering monophyletic taxa.
The basis for analysis is the cladogram. A cladogram is a dendrogram where the branch points are defined by synapomorphies. If absence
of a particular tooth cusp is a plesiomorphic character state and others have the cusp (cusp 1) present, the branch point would separate the two groups. 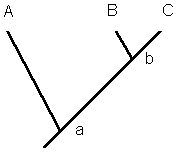 If some in the second group also have an additional cusp 2, then an additional separation would be shown. In the adjacent cladogram, the plesiomorphic
state (no cusp) is represented by the ventral stem; "A" retains this state. Cusp 1 is present in those forms beyond the node (branch point) and thus is a synapomorphy for "B" and "C". Cusp 2
represents the evolution of a second new cusp beyond the node of "B" and "C" and thus taxon "C" has both cusps 1 and 2, but taxon "B" has only cusp 1. Since taxon C has a unique character state
evolved since its last common ancestor (its ancestor shared with "B", Cusp 2 is an autapomorphy.
If some in the second group also have an additional cusp 2, then an additional separation would be shown. In the adjacent cladogram, the plesiomorphic
state (no cusp) is represented by the ventral stem; "A" retains this state. Cusp 1 is present in those forms beyond the node (branch point) and thus is a synapomorphy for "B" and "C". Cusp 2
represents the evolution of a second new cusp beyond the node of "B" and "C" and thus taxon "C" has both cusps 1 and 2, but taxon "B" has only cusp 1. Since taxon C has a unique character state
evolved since its last common ancestor (its ancestor shared with "B", Cusp 2 is an autapomorphy.
A cladogram is not necessarily a phylogenetic dendrogram; it merely shows the relationships in terms of monophyly ("B" and "C" are believed to form a monophyletic group because they share an apomorphy and that is evidence that they have inherited it from a common ancestor). It is quite possible to interpret a phylogenetic tree from the cladogram in which "B" is the ancestor of "C" (this still is monophyly since "B" and "C" would still share a common ancestor shared by no other taxon—namely, the immediate ancestor of "B" in this case).
To do an analysis, characters are chosen and character states determined for the taxa involved. Also, polarity must be established (that is, what is plesiomorphic and what is apomorphic in any series of character states). For the latter, often an outgroup is chosen—a higher-level, related taxon. Character states shared with the outgroup are presumed to be plesiomorphic. For example, if you were working with the family Felidae and used the Canidae as the outgroup, you would take the presence of the carnassial pair of teeth as being plesiomorphic since it's present in both taxa, indicating that it had arisen in a common ancestor of cats and dogs. As such, it would be of no value in trying to determine the relationships within the family Felidae.
Despite neat theory, there are (as in most scientific endeavors) problems in practice. A general problem is that posed by parallelisms, convergences, and reversals. These show up in inconsistencies (in
our example, as if "A" and "C" shared some synapomorphies while "B" and "C" shared others). Generally, the principle of parsimony is called upon; those relationships that require the fewest ad hoc
explanations are chosen as the best supported. 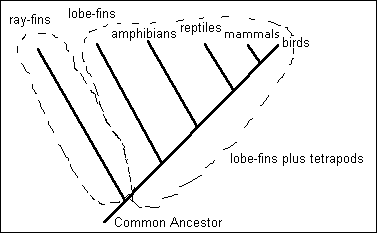
Other problems, particularly in paleontology, have to do with inadequate suites of characters and determining character-state polarity with long-extinct taxa.
From the point of view of traditional taxonomy, cladistics results in extreme vertical classification and, if used as a formal taxonomy, results in situations uncomfortable to many. A favorite example is that lobe-finned fishes are placed in a high-level taxon (with their descendents, the tetrapods) separate from the ray-finned fishes. Also, such paraphyletic groups as reptiles (Class Aves and Class Mammalia are not included within the Class Reptilia in traditional taxonomy) undergo major changes. Under one scenario, for example, the reptiles would originate at the branch point where birds originate, and of course would not include "reptilian" taxa that split off earlier than the Jurassic. Alternatively, the Reptilia could originate at the branch point where the mammals originate, and birds then would be a subdivision of the class Reptilia. Although various groups are working on taxonomic treatments, the situation is considered a continuing problem by many. This is one reason we are sticking mostly with the classical taxonomy in this class.
Last Update: 10 Jan 2008
Centennial Museum and Department of Biological Sciences, The University of Texas at El Paso